Changing the Linear Components¶
Here we calculate trends using the SAGE II/OSIRIS/OMPS-LP Dataset. Trends are calculated in the following ways
With two linear terms that have a join point at 1997.
Same as above but with the join point changed to 2000
By fitting ozone anomalies without any linear component, and then fitting the residuals to a linear term
Using two orthogonal forms of the EESC
Start with some imports and loading in the data
[2]:
import xarray as xr
import numpy as np
from LOTUS_regression.regression import regress_all_bins
from LOTUS_regression.predictors import load_data
import LOTUS_regression.predictors.download as download
import LOTUS_regression.plotting.trends as trends
import matplotlib.pyplot as plt
from datetime import datetime
import pandas as pd
import time
[3]:
MERGED_FILE = r'/home/runner/work/lotus-regression/lotus-regression/test_data/S2_OS_OMPS/MERGED_LOTUS.nc'
mzm_data = xr.open_dataset(MERGED_FILE, engine='netcdf4')
We have our default set of predictors which contains linear terms joining at 1997
[4]:
predictors = load_data('pred_baseline_pwlt.csv').to_period('M')
print(predictors.columns)
Index(['enso', 'solar', 'qboA', 'qboB', 'aod', 'linear_pre', 'linear_post',
'constant'],
dtype='object')
And our default set of trends
[5]:
results = regress_all_bins(predictors, mzm_data['relative_anomaly'], tolerance=0.1)
# Convert to ~ percent
results *= 100
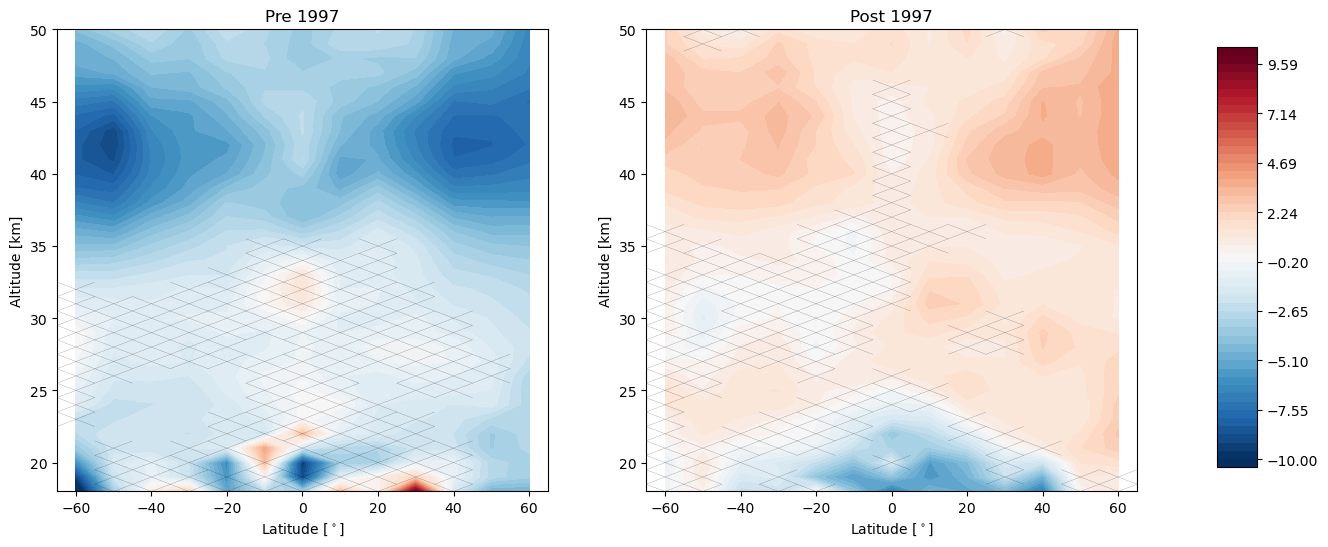
trends.pre_post_with_confidence(results, x='mean_latitude', y='altitude', ylim=(18, 50), log_y=False, figsize=(16, 6),
x_label='Latitude [$^\circ$]', y_label='Altitude [km]', pre_title='Pre 1997',
post_title='Post 1997')
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
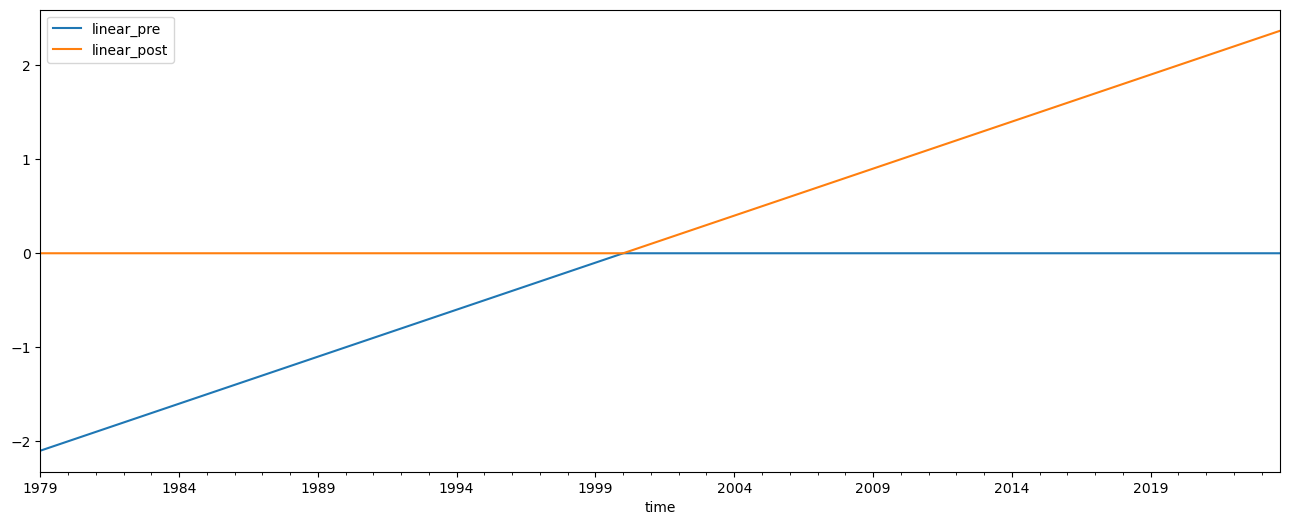
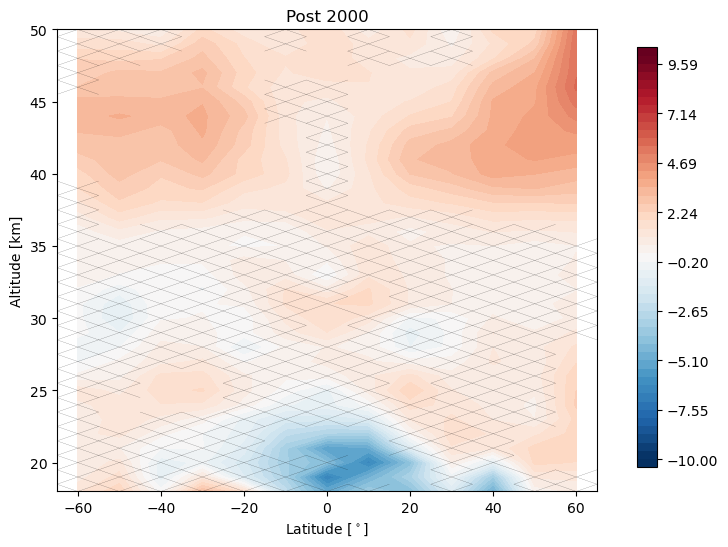
Now we change our inflection point in the linear trend to 2000 instead of 1997, and repeat the analysis
[6]:
predictors[['linear_pre', 'linear_post']] = download.load_linear(inflection=2000)[['pre','post']]
predictors[['linear_pre', 'linear_post']].plot(figsize=(16, 6))
[6]:
<Axes: xlabel='time'>
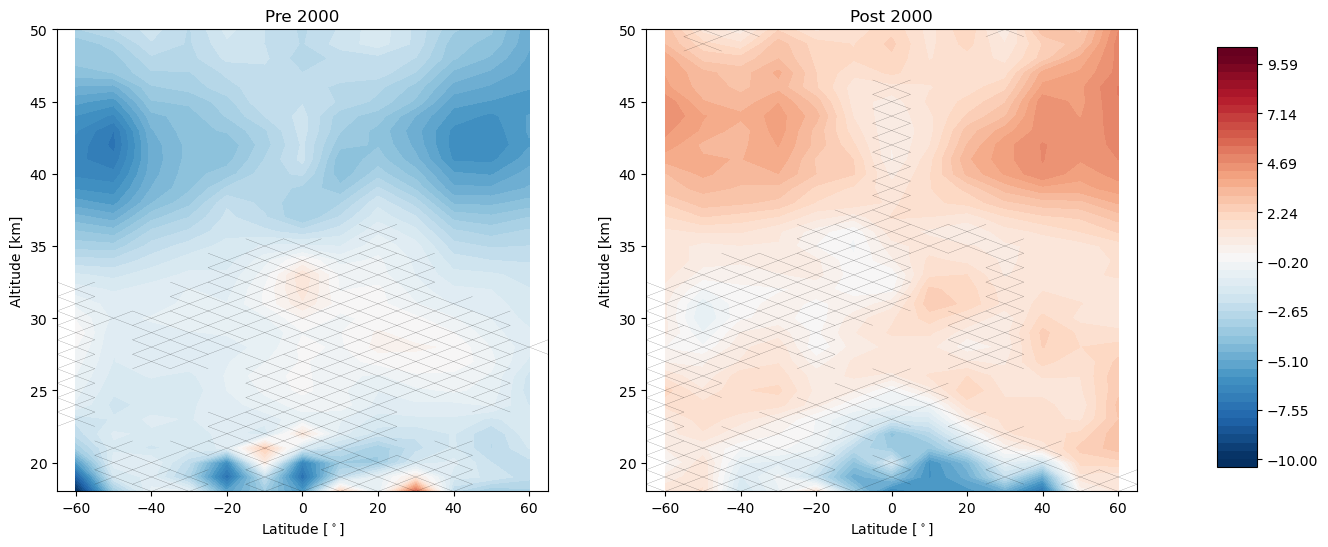
[7]:
results = regress_all_bins(predictors, mzm_data['relative_anomaly'], tolerance=0.1)
# Convert to ~ percent
results *= 100
trends.pre_post_with_confidence(results, x='mean_latitude', y='altitude', ylim=(18, 50), log_y=False, figsize=(16, 6),
x_label='Latitude [$^\circ$]', y_label='Altitude [km]', pre_title='Pre 2000',
post_title='Post 2000')
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Now we remove the linear terms from our predictors and tell the regression call to post fit a trend to the residuals after the year 2000
[8]:
predictors = predictors.drop(['linear_pre', 'linear_post'], axis=1)
results = regress_all_bins(predictors, mzm_data['relative_anomaly'], tolerance=0.1, post_fit_trend_start='2000-01-01')
# Convert to ~ percent
results *= 100
trends.post_with_confidence(results, x='mean_latitude', y='altitude', ylim=(18, 50), log_y=False, figsize=(8, 6),
x_label='Latitude [$^\circ$]', y_label='Altitude [km]',
post_title='Post 2000')
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
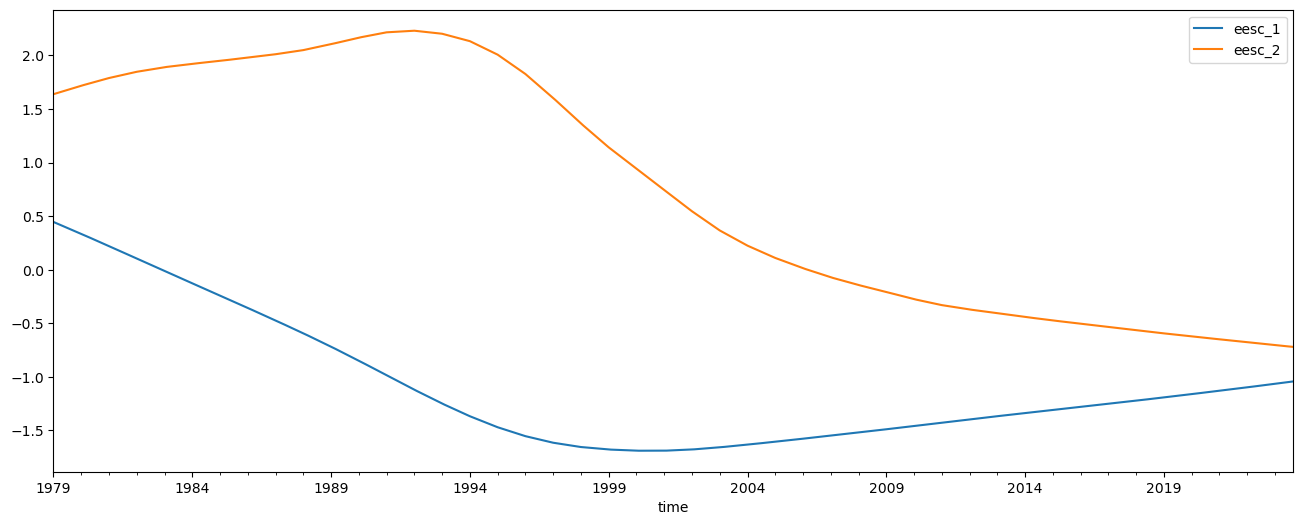
Lastly we add two orthogonal EESC terms and redo the regression
[9]:
predictors[['eesc_1', 'eesc_2']] = download.load_orthogonal_eesc('/home/runner/work/lotus-regression/lotus-regression/test_data/EESC.txt')[['eesc_1', 'eesc_2']]
predictors[['eesc_1', 'eesc_2']].plot(figsize=(16, 6))
results = regress_all_bins(predictors, mzm_data['relative_anomaly'], tolerance=0.1)
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
Intel MKL ERROR: Parameter 5 was incorrect on entry to DTRTRI.
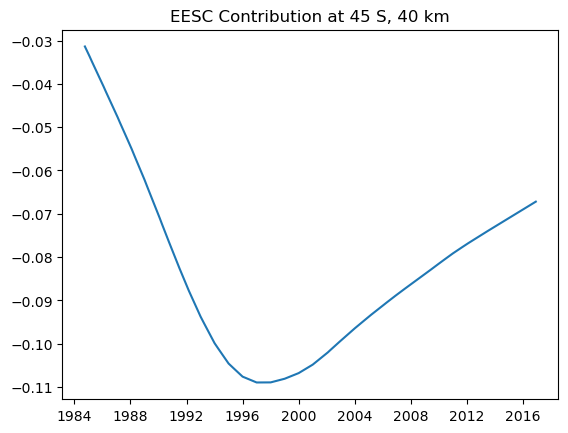
The results are trickier to interpret, but we can combine the two EESC terms to find the total contribution in each bin for the EESC
[10]:
predictors = predictors[(pd.to_datetime(predictors.index.to_timestamp()) >= mzm_data.time.values[0]) & (pd.to_datetime(predictors.index.to_timestamp()) <= mzm_data.time.values[-1])]
eesc_contrib = results['eesc_1'].values[:, :, np.newaxis] * predictors['eesc_1'].values[np.newaxis, np.newaxis, :] +\
results['eesc_2'].values[:, :, np.newaxis] * predictors['eesc_2'].values[np.newaxis, np.newaxis, :]
plt.plot(mzm_data.time.values, eesc_contrib[2, 40, :].T)
np.shape(eesc_contrib)
plt.title('EESC Contribution at 45 S, 40 km')
[10]:
Text(0.5, 1.0, 'EESC Contribution at 45 S, 40 km')
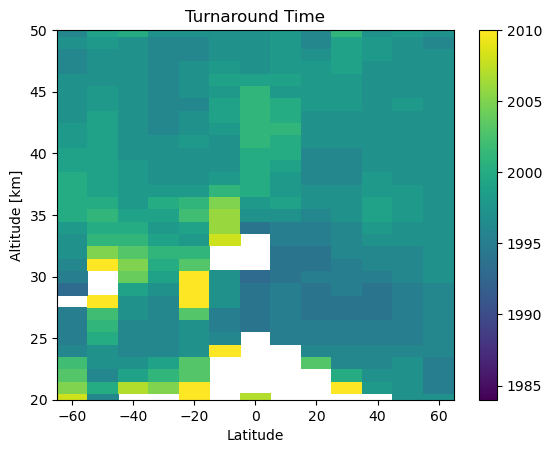
With some annoying time conversions we can also find the predicted turnaround time in each bin
[11]:
turnaround_index = np.argmin(eesc_contrib, axis=2)
turnaround_times = mzm_data.time.values[turnaround_index]
def to_year_fraction(date):
def since_epoch(date): # returns seconds since epoch
return time.mktime(date.timetuple())
try:
ts = (np.datetime64(date, 'ns') - np.datetime64('1970-01-01T00:00:00Z')) / np.timedelta64(1, 's')
except:
print(np.datetime64(date))
return np.nan
dt = datetime.utcfromtimestamp(ts)
s = since_epoch
year = dt.year
startOfThisYear = datetime(year=year, month=1, day=1)
startOfNextYear = datetime(year=year+1, month=1, day=1)
yearElapsed = s(dt) - s(startOfThisYear)
yearDuration = s(startOfNextYear) - s(startOfThisYear)
fraction = yearElapsed / yearDuration
return dt.year + fraction
turnaround_times = np.frompyfunc(to_year_fraction, 1, 1)(turnaround_times).astype(float)
turnaround_times[turnaround_index == 0] = np.nan
turnaround_times[turnaround_index == len(mzm_data.time.values)-1] = np.nan
plt.pcolor(mzm_data.mean_latitude.values, mzm_data.altitude.values, np.ma.masked_invalid(turnaround_times.T))
plt.clim(1984, 2010)
plt.ylim(20, 50)
plt.ylabel('Altitude [km]')
plt.xlabel('Latitude')
plt.title('Turnaround Time')
plt.colorbar()
[11]:
<matplotlib.colorbar.Colorbar at 0x7f6911705210>